Overview
This notebook covers a implementation of a basic neural network. It shows initalizing weights and set up an architecture
Code
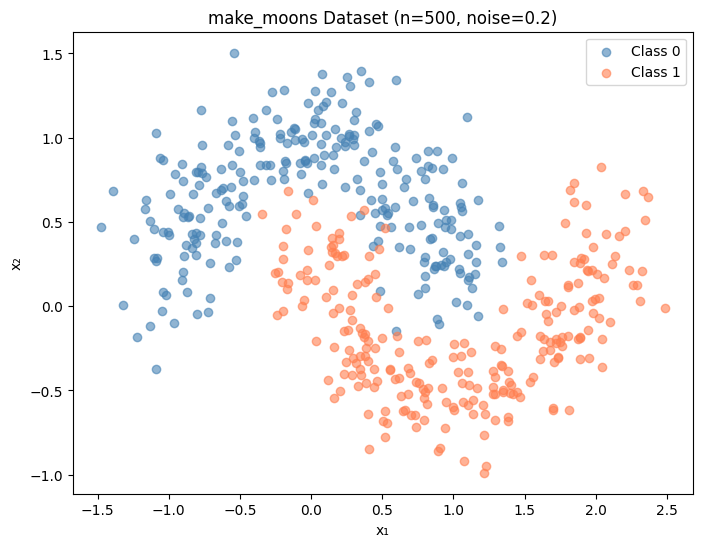
import numpy as npimport matplotlib.pyplot as pltfrom sklearn.datasets import make_moonsfrom sklearn.tree import DecisionTreeClassifier# Generate the dataset = make_moons(n_samples= 500 , noise= 0.2 , random_state= 42 )= (8 , 6 ))== 0 , 0 ], X[y== 0 , 1 ], color= 'steelblue' , label= 'Class 0' , alpha= 0.6 )== 1 , 0 ], X[y == 1 , 1 ], c= 'coral' , label= 'Class 1' , alpha= 0.6 )'make_moons Dataset (n=500, noise=0.2)' )'x₁' )'x₂' )
Code
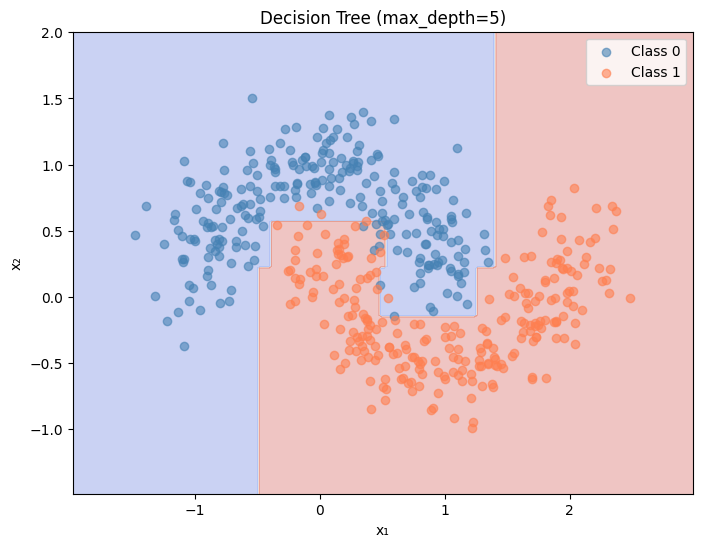
def plot_decision_boundary(model_predict, X, y, title, ax= None ):"""Plot the decision boundary of a classifier.""" if ax is None := plt.subplots(figsize= (8 , 6 ))= X[:, 0 ].min () - 0.5 , X[:, 0 ].max () + 0.5 = X[:, 1 ].min () - 0.5 , X[:, 1 ].max () + 0.5 = np.meshgrid(np.linspace(x_min, x_max, 200 ),200 ))= np.c_[xx.ravel(), yy.ravel()]= model_predict(grid).reshape(xx.shape)= 0.3 , cmap= 'coolwarm' )== 0 , 0 ], X[y == 0 , 1 ], c= 'steelblue' , label= 'Class 0' , alpha= 0.6 )== 1 , 0 ], X[y == 1 , 1 ], c= 'coral' , label= 'Class 1' , alpha= 0.6 )'x₁' )'x₂' )# Train and plot decision tree = DecisionTreeClassifier(max_depth= 5 , random_state= 42 )'Decision Tree (max_depth=5)' )
Code
def sigmoid(z):"""Sigmoid activation function.""" return 1 / (1 + np.exp(- z))def sigmoid_derivative(a):"""Derivative of sigmoid, given the sigmoid output a = sigmoid(z).""" return a * (1 - a)
Code
= np.array([- 5 , - 1 , 0 , 1 , 5 ])print (f"sigmoid( { z} ) = { sigmoid(z). round (4 )} " )# Expected: [0.0067, 0.2689, 0.5, 0.7311, 0.9933]
sigmoid([-5 -1 0 1 5]) = [0.0067 0.2689 0.5 0.7311 0.9933]
Code
42 )# Shapes: (rows = inputs from previous layer, cols = neurons in this layer) = np.random.randn(2 , 8 ) * 0.5 # (2 inputs, 8 hidden) → shape (2, 8) = np.zeros((1 , 8 )) # (1, 8) = np.random.randn(8 , 1 ) * 0.5 # (8 hidden, 1 output) → shape (8, 1) = np.zeros((1 , 1 )) # (1, 1) print (f"W1 shape: { W1. shape} " ) # (2, 8) print (f"b1 shape: { b1. shape} " ) # (1, 8) print (f"W2 shape: { W2. shape} " ) # (8, 1) print (f"b2 shape: { b2. shape} " ) # (1, 1)
W1 shape: (2, 8)
b1 shape: (1, 8)
W2 shape: (8, 1)
b2 shape: (1, 1)
Code
def forward(X, W1, b1, W2, b2):""" Forward pass through the network. X shape: (n_samples, 2) Z1 shape: (n_samples, 8) — hidden layer pre-activation A1 shape: (n_samples, 8) — hidden layer post-activation Z2 shape: (n_samples, 1) — output pre-activation A2 shape: (n_samples, 1) — output (prediction) """ = X @ W1 + b1 # (n, 2) @ (2, 8) + (1, 8) = (n, 8) = sigmoid(Z1) # (n, 8) = A1 @ W2 + b2 # (n, 8) @ (8, 1) + (1, 1) = (n, 1) = sigmoid(Z2) # (n, 1) return Z1, A1, Z2, A2# Test with a few samples = forward(X[:5 ], W1, b1, W2, b2)print (f"Predictions for first 5 samples: { A2. flatten(). round (4 )} " )print (f"Actual labels: { y[:5 ]} " )
Predictions for first 5 samples: [0.2532 0.291 0.2352 0.368 0.3455]
Actual labels: [1 0 1 0 0]
Code
def compute_loss(y_true, y_pred):"""Binary cross-entropy loss.""" = y_true.shape[0 ]# Clip predictions to avoid log(0) = np.clip(y_pred, 1e-8 , 1 - 1e-8 )= - np.mean(y_true * np.log(y_pred) + (1 - y_true) * np.log(1 - y_pred))return loss
Code
def backward(X, y_true, Z1, A1, Z2, A2, W1, b1, W2, b2):""" Backward pass — compute gradients for all weights and biases. Returns gradients: dW1, db1, dW2, db2 """ = X.shape[0 ]= y_true.reshape(- 1 , 1 ) # (n, 1) # Output layer error = A2 - y_true # (n, 1) — how wrong is each prediction? # Gradients for W2, b2 = (A1.T @ dZ2) / n # (8, 1) — average gradient across samples = np.sum (dZ2, axis= 0 , keepdims= True ) / n # (1, 1) # Propagate error back to hidden layer = dZ2 @ W2.T # (n, 8) = dA1 * sigmoid_derivative(A1) # (n, 8) — chain rule with activation # Gradients for W1, b1 = (X.T @ dZ1) / n # (2, 8) = np.sum (dZ1, axis= 0 , keepdims= True ) / n # (1, 8) return dW1, db1, dW2, db2
Code
# Hyperparameters = 0.5 = 1000 # Re-initialize weights 42 )= np.random.randn(2 , 8 ) * 0.5 = np.zeros((1 , 8 ))= np.random.randn(8 , 1 ) * 0.5 = np.zeros((1 , 1 ))# Track loss for plotting = []for epoch in range (epochs):# Forward = forward(X, W1, b1, W2, b2)# Loss = compute_loss(y, A2)# Backward = backward(X, y, Z1, A1, Z2, A2, W1, b1, W2, b2)# Update weights (gradient descent) -= learning_rate * dW1-= learning_rate * db1-= learning_rate * dW2-= learning_rate * db2if epoch % 100 == 0 := np.mean((A2.flatten() > 0.5 ) == y)print (f"Epoch { epoch:4d} | Loss: { loss:.4f} | Accuracy: { accuracy:.3f} " )print (f" \n Final — Loss: { losses[- 1 ]:.4f} | Accuracy: { np. mean((A2.flatten() > 0.5 ) == y):.3f} " )
Epoch 0 | Loss: 0.7948 | Accuracy: 0.500
Epoch 100 | Loss: 1.0295 | Accuracy: 0.846
Epoch 200 | Loss: 1.2687 | Accuracy: 0.860
Epoch 300 | Loss: 1.3746 | Accuracy: 0.862
Epoch 400 | Loss: 1.4233 | Accuracy: 0.860
Epoch 500 | Loss: 1.4460 | Accuracy: 0.860
Epoch 600 | Loss: 1.4573 | Accuracy: 0.860
Epoch 700 | Loss: 1.4636 | Accuracy: 0.860
Epoch 800 | Loss: 1.4678 | Accuracy: 0.860
Epoch 900 | Loss: 1.4710 | Accuracy: 0.860
Final — Loss: 1.4737 | Accuracy: 0.860
Code
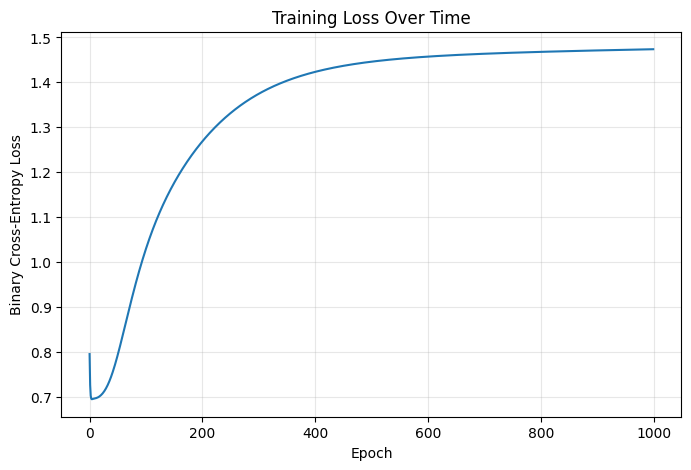
= (8 , 5 ))'Training Loss Over Time' )'Epoch' )'Binary Cross-Entropy Loss' )True , alpha= 0.3 )
Code
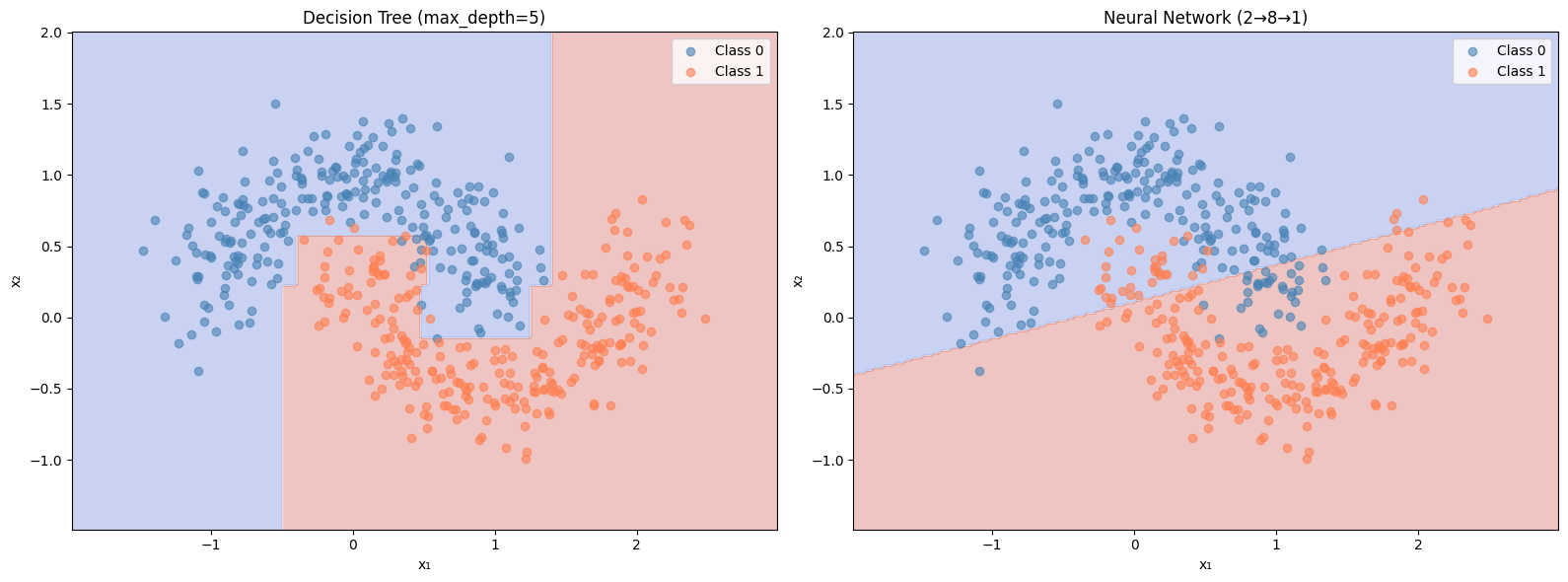
def nn_predict(X_input):"""Predict using our trained neural network.""" = forward(X_input, W1, b1, W2, b2)return (A2.flatten() > 0.5 ).astype(int )= plt.subplots(1 , 2 , figsize= (16 , 6 ))# Decision tree 'Decision Tree (max_depth=5)' , ax= axes[0 ])# Neural network 'Neural Network (2→8→1)' , ax= axes[1 ])
#PyTorch implementation of the same neural network import torch import torch.nn as nn import numpy as np import matplotlib.pyplot as plt
— Convert data to PyTorch tensors —
X_tensor = torch.FloatTensor(X) y_tensor = torch.FloatTensor(y).reshape(-1, 1)
— Initialize model —
torch.manual_seed(42) model = NeuralNet()
Match your original weight scale of 0.5
with torch.no_grad(): model.layer1.weight.data = 0.5 model.layer2.weight.data = 0.5
— Hyperparameters —
criterion = nn.BCELoss() # same loss as compute_loss() optimizer = torch.optim.SGD(model.parameters(), lr=0.5) # same lr as yours
— Training loop —
losses = []
for epoch in range(1000): # Forward pass A2 = model(X_tensor)
# Loss
loss = criterion(A2, y_tensor)
losses.append(loss.item())
# Backward pass
optimizer.zero_grad()
loss.backward()
optimizer.step()
if epoch % 100 == 0:
preds = (A2.detach() > 0.5).float()
accuracy = (preds == y_tensor).float().mean()
print(f"Epoch {epoch:4d} | Loss: {loss:.4f} | Accuracy: {accuracy:.3f}")print(f”— Loss: {losses[-1]:.4f} | Accuracy: {(((model(X_tensor).detach() > 0.5).float() == y_tensor).float().mean()):.3f}“)
— Plot loss curve —
plt.plot(losses) plt.xlabel(“Epoch”) plt.ylabel(“Loss”) plt.title(“Training Loss”) plt.show()
#PyTorch implementation of GNN
import torch import torch.nn.functional as F from torch_geometric.datasets import Planetoid from torch_geometric.nn import GCNConv
— Dataset —
dataset = Planetoid(root=“/tmp/pyg_data”, name=“Cora”) data = dataset[0] device = torch.device(“cuda” if torch.cuda.is_available() else “cpu”) data = data.to(device)
— Model —
class GCN(torch.nn.Module): def init (self, in_channels: int, hidden_channels: int, out_channels: int): super().__init__() self.conv1 = GCNConv(in_channels, hidden_channels) self.conv2 = GCNConv(hidden_channels, out_channels)
def forward(self, x, edge_index):
x = self.conv1(x, edge_index)
x = F.relu(x)
x = F.dropout(x, p=0.5, training=self.training)
x = self.conv2(x, edge_index)
return xmodel = GCN(dataset.num_node_features, 16, dataset.num_classes).to(device) optimizer = torch.optim.Adam(model.parameters(), lr=0.01, weight_decay=5e-4)
— Training —
model.train() for epoch in range(1, 201): optimizer.zero_grad() out = model(data.x, data.edge_index) loss = F.cross_entropy(out[data.train_mask], data.y[data.train_mask]) loss.backward() optimizer.step()
if epoch % 50 == 0:
model.eval()
with torch.no_grad():
pred = model(data.x, data.edge_index).argmax(dim=1)
correct = (pred[data.test_mask] == data.y[data.test_mask]).sum()
acc = correct / data.test_mask.sum()
print(f"Epoch {epoch:>3d} Loss: {loss:.4f} Test Acc: {acc:.4f}")
model.train()